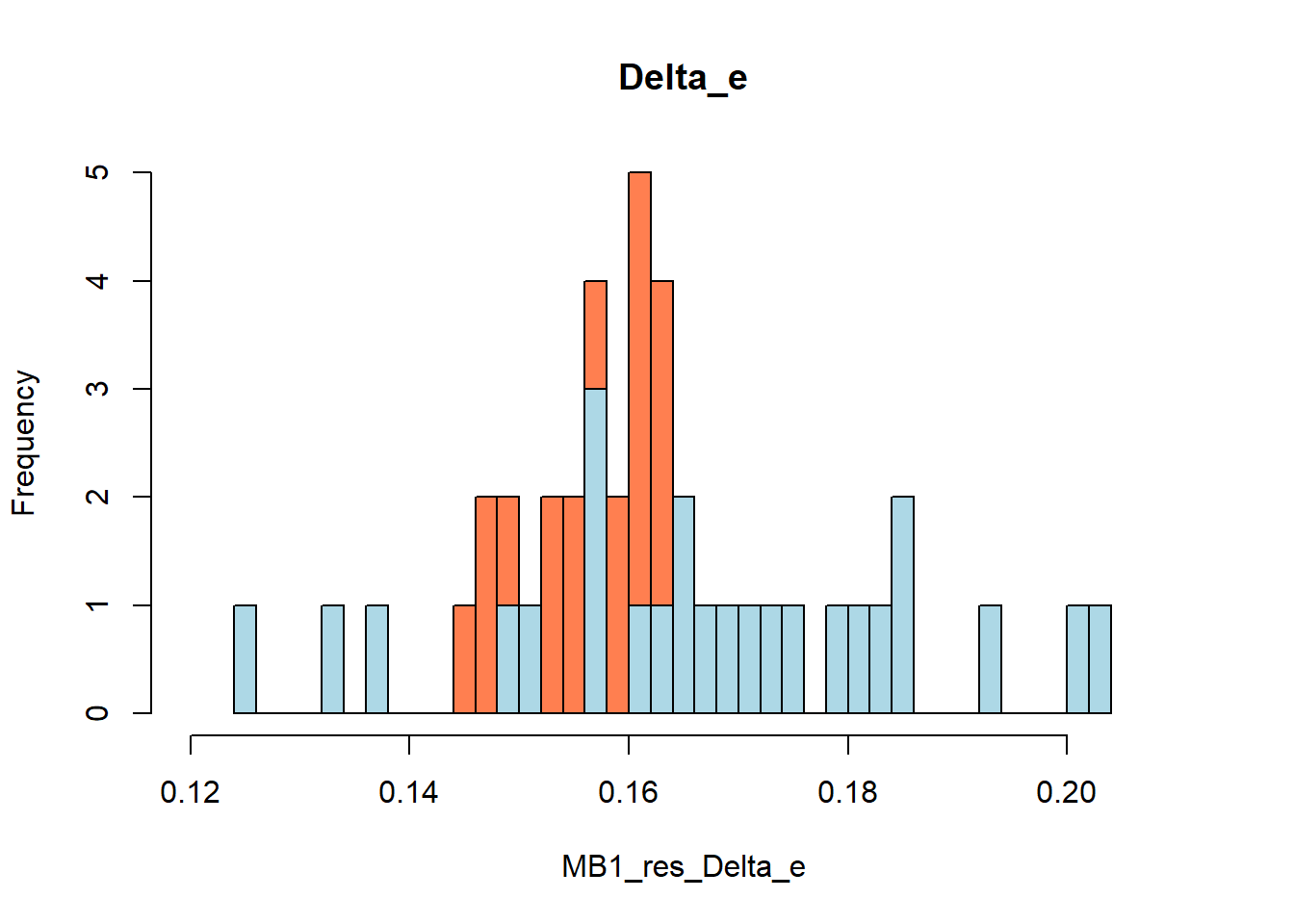
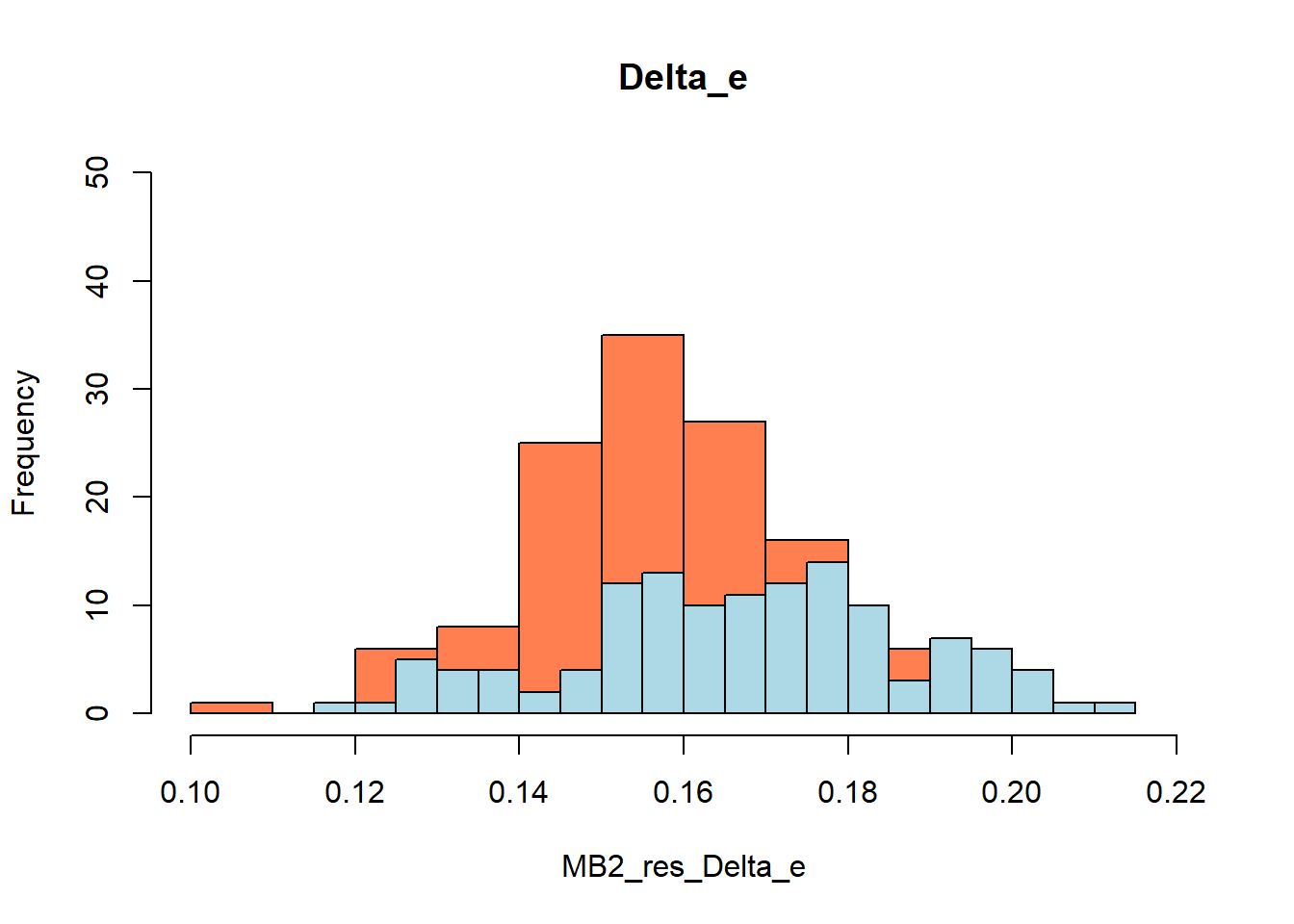
#function and needed packages to generate bootstrap estimates for mean QALY/TC and incremental
#differences when running a OLS/SUR model after applying MICE to handle missing data
#note: different ways to combine MI and bootstrapping estimates can be obtained based on the inputs provided
library(systemfit)
library(data.table)
library(rlang)
library(bootstrap)
library(emmeans)
library(mice)
boot_mi_ec <- function(x, B, QALYreg, TCreg, method = "OLS",
profile_QALY="default", profile_TC="default",
trt_pos = 2, combine="MB"){
#the following lines are needed to make sure proper inputs are given
if(!inherits(x, c("mids"))) {stop("Only objects of class 'mids' can be used")}
if(!is.data.frame(x$data)){stop("data needs to be a data frame object")}
if(!is.numeric(B)){stop("please provide number of bootstrap iterations")}
if(B<=0 | !B%%1==0){stop("please provide number of bootstrap iterations")}
if(!is_formula(QALYreg)){stop("please provide formula for QALY model")}
if(!is_formula(TCreg)){stop("please provide formula for TC model")}
if(!method %in% c("OLS","SUR")){stop("please provide valid method name")}
if(!combine %in% c("MB","BM")){stop("please provide valid combination name")}
if(!is.numeric(trt_pos) | length(trt_pos)!=1 | trt_pos<=0){stop("please provide valid trt indicator position in regressions")}
data <- x$data #original data set
n <- dim(data)[1] #original sample size
#n covariates
nX_e <- dim(model.matrix(QALYreg, data))[2]
nX_c <- dim(model.matrix(TCreg, data))[2]
#extract name of trt indicator and outcomes from provided formula
trt_name_e <- all.vars(QALYreg)[trt_pos]
trt_name_c <- all.vars(TCreg)[trt_pos]
if(trt_name_e != trt_name_c){stop("please provide same trt variable name and position in QALY and TC formuale")}
QALY_name <- all.vars(QALYreg)[1]
TC_name <- all.vars(TCreg)[1]
#check if trt indicator is factor and store its levels
if(is.factor(data[,trt_name_e])){
trt_fact <- TRUE
trt_lev <- levels(data[,trt_name_e])} else {
trt_fact <- FALSE
trt_lev <- unique(data[,trt_name_e])}
if(length(trt_lev)!=2){stop("The function only allows comparison between two trt groups")}
#check that correct profile provided or set default
if(profile_QALY != "default"){
if(!is.vector(profile_QALY) | length(profile_QALY)!=nX_e){stop("provide valid profile for QALYreg")}}
if(profile_TC != "default"){
if(!is.vector(profile_TC) | length(profile_TC)!=nX_c){stop("provide valid profile for TCreg")}}
#extract MI infor from input object
M <- x$m #n imputations
pm <- x$predictorMatrix #predictor matrix
#remove trt as predictor
pm[,trt_name_e] <- 0
#pm <- pm[-which(colnames(pm)==trt_name_e),-which(colnames(pm)==trt_name_e)]
meth <- x$method #imputation methods
niter <- x$iteration #n iterations before sampling
#split original data by treatment group
x_trt1 <- data.frame(x$data[x$data[,trt_name_e]==unique(data[,trt_name_e])[1],])
x_trt2 <- data.frame(x$data[x$data[,trt_name_e]==unique(data[,trt_name_e])[2],])
#generate imputations and bootstrap estimates based on inputs
if(combine == "MB"){ #outer loop=MI + inner loop=Boot
#prepare empty objects to contain bootstrapped estimates
data_MB_list <- list()
coeff_c <- coeff_e <- matrix(NA, nrow = M, ncol = B)
em_c_ctr <- em_e_ctr <- matrix(NA, nrow = M, ncol = B)
em_c_int <- em_e_int <- matrix(NA, nrow = M, ncol = B)
data_imp <- list()
for(m in 1:M){
#apply MI to each arm based on arguments from mice objects provided as input
mice_data_trt1 <- mice(x_trt1, predictorMatrix = pm,
method=meth,m = M, maxit = niter, print = FALSE)
mice_data_trt2 <- mice(x_trt2, predictorMatrix = pm,
method=meth,m = M, maxit = niter, print = FALSE)
#combine the imputed datasets across groups
data_imp <- rbind(mice_data_trt1, mice_data_trt2)
#extract imputed data sets
data_imp[[m]] <- data.table(complete(data_imp,m))
for(i in 1:B){
#sample with replacement for each imputed data set
data_MB_list[[i]] <- data_imp[[m]][sample(.N, n, replace = T)]
#fit model
model_MB_ec <- systemfit(list(QALYreg = QALYreg, TCreg = TCreg),
method=method, data=data_MB_list[[i]])
#extract covariate values
X_e <- model.matrix(model_MB_ec$eq[[1]])
X_c <- model.matrix(model_MB_ec$eq[[2]])
#define QALYreg profile
if(profile_QALY == "default"){
profile_b_QALY <- apply(X_e, 2, mean, na.rm=T)
} else {profile_b_QALY <- profile_QALY}
profile_b_QALY_ctr <- profile_b_QALY_int <- profile_b_QALY
profile_b_QALY_ctr[trt_pos] <- 0 #set profile for comparator
profile_b_QALY_int[trt_pos] <- 1 #set profile for reference
#define TCreg profile
if(profile_TC == "default"){
profile_b_TC <- apply(X_c, 2, mean, na.rm=T)
} else {profile_b_TC <- profile_TC}
profile_b_TC_ctr <- profile_b_TC_int <- profile_b_TC
profile_b_TC_ctr[trt_pos] <- 0 #set profile for comparator
profile_b_TC_int[trt_pos] <- 1 #set profile for reference
#extract coefficient estimates from each model
coeff_e[m,i] <- summary(model_MB_ec$eq[[1]])$coefficients[trt_pos,"Estimate"]
coeff_c[m,i] <- summary(model_MB_ec$eq[[2]])$coefficients[trt_pos,"Estimate"]
#compute linear combination of parameters
em_e_ctr[m,i] <- t(profile_b_QALY_ctr) %*% summary(model_MB_ec$eq[[1]])$coefficients[,"Estimate"]
em_e_int[m,i] <- t(profile_b_QALY_int) %*% summary(model_MB_ec$eq[[1]])$coefficients[,"Estimate"]
em_c_ctr[m,i] <- t(profile_b_TC_ctr) %*% summary(model_MB_ec$eq[[2]])$coefficients[,"Estimate"]
em_c_int[m,i] <- t(profile_b_TC_int) %*% summary(model_MB_ec$eq[[2]])$coefficients[,"Estimate"]
}
}
}
if(combine == "BM"){ #outer loop=Boot + inner loop=MI
#prepare empty objects to contain bootstrapped estimates
data_BM_list <- list()
coeff_c <- coeff_e <- matrix(NA, nrow = B, ncol = M)
em_c_ctr <- em_e_ctr <- matrix(NA, nrow = B, ncol = M)
em_c_int <- em_e_int <- matrix(NA, nrow = B, ncol = M)
data.dt <- data.table(x$data)
data_imp <- list()
for(i in 1:B){
#sample with replacement from original data set
data_BM_list[[i]] <- as.data.frame(data.dt[sample(.N, n, replace = T)])
#split data by arm
data_BM_trt1 <- data.frame(data_BM_list[[i]][data_BM_list[[i]][,trt_name_e]==unique(x$data[,trt_name_e])[1],])
data_BM_trt2 <- data.frame(data_BM_list[[i]][data_BM_list[[i]][,trt_name_e]==unique(x$data[,trt_name_e])[2],])
#apply MI per arm based on arguments from mice objects provided as input
mice_data_boot_trt1 <- mice(data_BM_trt1, predictorMatrix = pm,
method=meth,m = M, maxit = niter, print = FALSE)
mice_data_boot_trt2 <- mice(data_BM_trt2, predictorMatrix = pm,
method=meth,m = M, maxit = niter, print = FALSE)
#combine the imputed datasets across groups
mice_data_boot <- rbind(mice_data_boot_trt1, mice_data_boot_trt2)
for(m in 1:M){
#extract each imputed data set
data_imp[[m]] <- complete(mice_data_boot, m)
#fit model to each imputed data set
model_BM_ec <- systemfit(list(QALYreg = QALYreg, TCreg = TCreg),
method=method, data=data_imp[[m]])
#extract covariate values
X_e <- model.matrix(model_BM_ec$eq[[1]])
X_c <- model.matrix(model_BM_ec$eq[[2]])
#define QALYreg profile
if(profile_QALY == "default"){
profile_b_QALY <- apply(X_e, 2, mean, na.rm=T)
} else {profile_b_QALY <- profile_QALY}
profile_b_QALY_ctr <- profile_b_QALY_int <- profile_b_QALY
profile_b_QALY_ctr[trt_pos] <- 0 #set profile for comparator
profile_b_QALY_int[trt_pos] <- 1 #set profile for reference
#define TCreg profile
if(profile_TC == "default"){
profile_b_TC <- apply(X_c, 2, mean, na.rm=T)
} else {profile_b_TC <- profile_TC}
profile_b_TC_ctr <- profile_b_TC_int <- profile_b_TC
profile_b_TC_ctr[trt_pos] <- 0 #set profile for comparator
profile_b_TC_int[trt_pos] <- 1 #set profile for reference
#extract coefficient estimates from each model
coeff_e[i,m] <- summary(model_BM_ec$eq[[1]])$coefficients[trt_pos,"Estimate"]
coeff_c[i,m] <- summary(model_BM_ec$eq[[2]])$coefficients[trt_pos,"Estimate"]
#compute linear combination of parameters
em_e_ctr[i,m] <- t(profile_b_QALY_ctr) %*% summary(model_BM_ec$eq[[1]])$coefficients[,"Estimate"]
em_e_int[i,m] <- t(profile_b_QALY_int) %*% summary(model_BM_ec$eq[[1]])$coefficients[,"Estimate"]
em_c_ctr[i,m] <- t(profile_b_TC_ctr) %*% summary(model_BM_ec$eq[[2]])$coefficients[,"Estimate"]
em_c_int[i,m] <- t(profile_b_TC_int) %*% summary(model_BM_ec$eq[[2]])$coefficients[,"Estimate"]
}
}
}
#create list objects to store all results
res_e_b_list <-list("Delta_e"=coeff_e,"mu_e_ctr"=em_e_ctr,"mu_e_int"=em_e_int)
res_c_b_list <-list("Delta_c"=coeff_c,"mu_c_ctr"=em_c_ctr,"mu_c_int"=em_c_int)
input_list <- list("data"=x$data,"method"=method, "trt_pos"=trt_pos, "QALYreg"=QALYreg,
"TCreg"=TCreg,"profile_QALY_ctr"=profile_b_QALY_ctr,
"profile_QALY_int"=profile_b_QALY_int,"profile_TC_ctr"=profile_b_TC_ctr,
"profile_TC_int"=profile_b_TC_int, "combine"=combine)
#compute overall list and return it as output from the function
res_ec_b_list <- list("QALY_boot"=res_e_b_list,"TC_boot"=res_c_b_list,"inputs"=input_list)
class(res_ec_b_list) <- "bootMICE"
return(res_ec_b_list)
}